The study was published in the prestigious journal Nucleic Acids Research and desribes a new small protein called CrsL. CrsL enables mycobacteria to adapt to temperature changes. It controls the switching on and off of genes that help mycobacteria grow at higher temperatures. The article was written in collaboration with research teams from the Institute of Microbiology of the Czech Academy of Sciences and the Faculty of Science of Charles University – groups focusing on microbial genetics and gene expression. It also involved scientists from CEITEC Masaryk University, whose expertise lies in the field of structural biology, and from the National Reference Laboratory for Mycobacteria at the National Institute of Public Health, which focuses on mycobacterial infections in the Czech Republic.
Mycobacteria include the slow-growing Mycobacterium tuberculosis, the bacterium responsible for tuberculosis, but also 200 other species of non-tuberculous bacteria. Previously, non-tuberculous mycobacteria were generally considered harmless, but in recent years the number of infections caused by these bacteria has been growing. They can cause problems especially in people with compromised immune systems, but in some cases also in otherwise healthy individuals. Many non-tuberculous mycobacteria can survive, for example, in shower heads, where they are exposed to fluctuations in temperature and humidity.
“We were interested in how mycobacteria adapt to changes. In the laboratory, we worked with non-pathogenic Mycobacterium smegmatis, which is used as a model bacterium not only for M. tuberculosis, but also for all other species of the genus Mycobacterium,” says Jarmila Hnilicová, who started the project at the Institute of Microbiology CAS and recently moved with her team to the Faculty of Science of Charles University.
Regulation of bacterial transcription, whereby information is transcribed from a DNA molecule to an RNA molecule, is essential for adaptation and survival. Bacterial polymerase, which “reads” genes on DNA, is responsible for transcription. Under stress, such as a change in temperature, polymerase preferentially reads the genes that help the bacteria survive in the given conditions.
“We were looking for proteins that are capable of binding to mycobacterial polymerase and influencing how it reads DNA. And we found a very small protein that probably no one had noticed before precisely because it is so small. However, when we remove it from the bacteria, the bacteria cannot grow at higher temperatures. And that’s interesting because mycobacteria, like their better-known neighbors Legionella, are able to survive in water pipes and are not bothered by higher temperatures. We know that mycobacteria are found in showers; according to some studies, they are even the most common group of bacteria in shower heads. That is why we want to know which genes help them survive in this environment,” says Jarmila Hnilicová. “This research began several years ago at IMIC CAS, and many colleagues from this institute have contributed significantly to it.”
Understanding the mechanisms that allow mycobacteria to survive in various environments, including homes and hospitals, will enable us to reduce the risk of infection in the future. The more we know about our tiny mycobacterial neighbors from showerheads, the better we will be able to defend ourselves against them.
PUBLICATION
Shoman et al., Expanding the CarD interaction network: CrsL is a novel transcription regulator in actinobacteria, Nucleic Acids Res, 2025 Nov 26;53(22):gkaf1342.

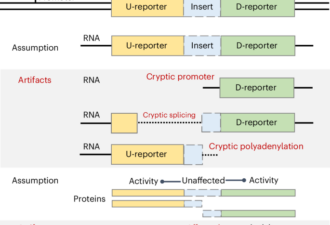
Graphical abstract – Shoman et al., Expanding the CarD interaction network: CrsL is a novel transcription regulator in actinobacteria, Nucleic Acids Res, 2025 Nov 26;53(22):gkaf1342.